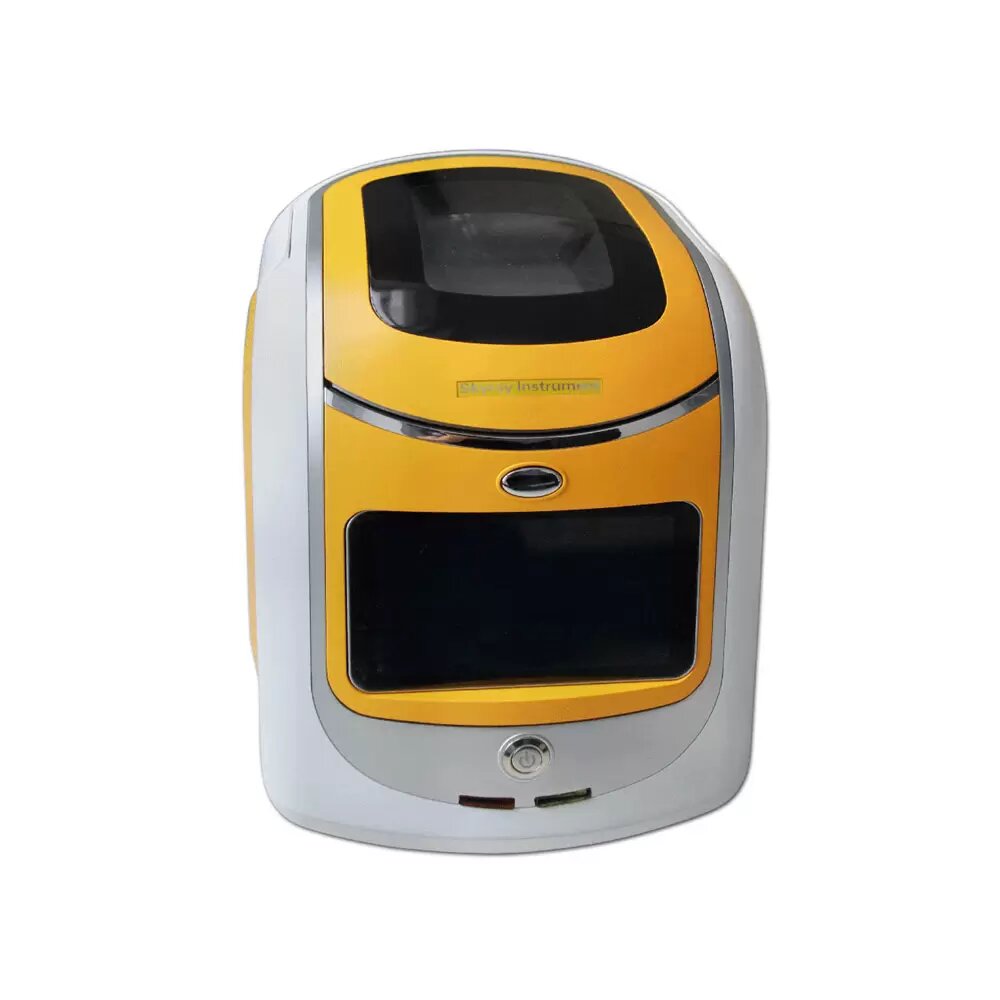
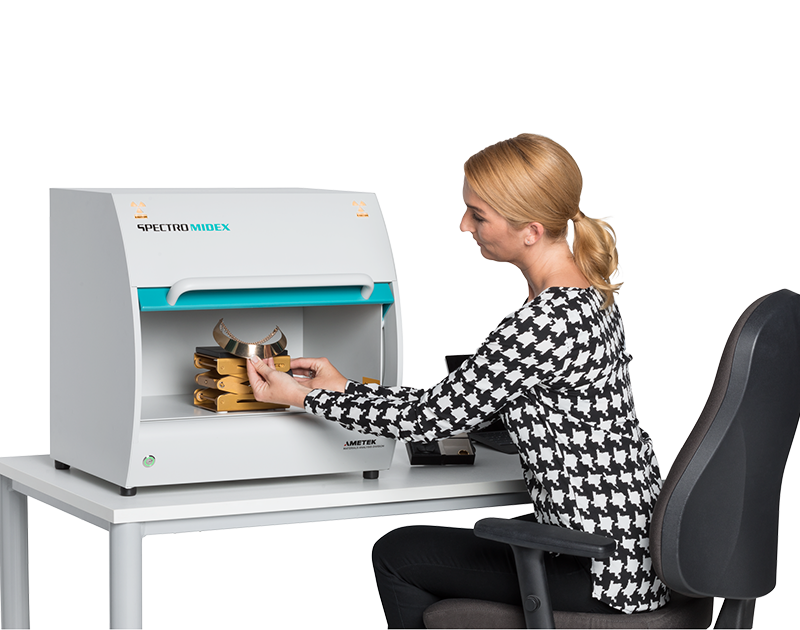
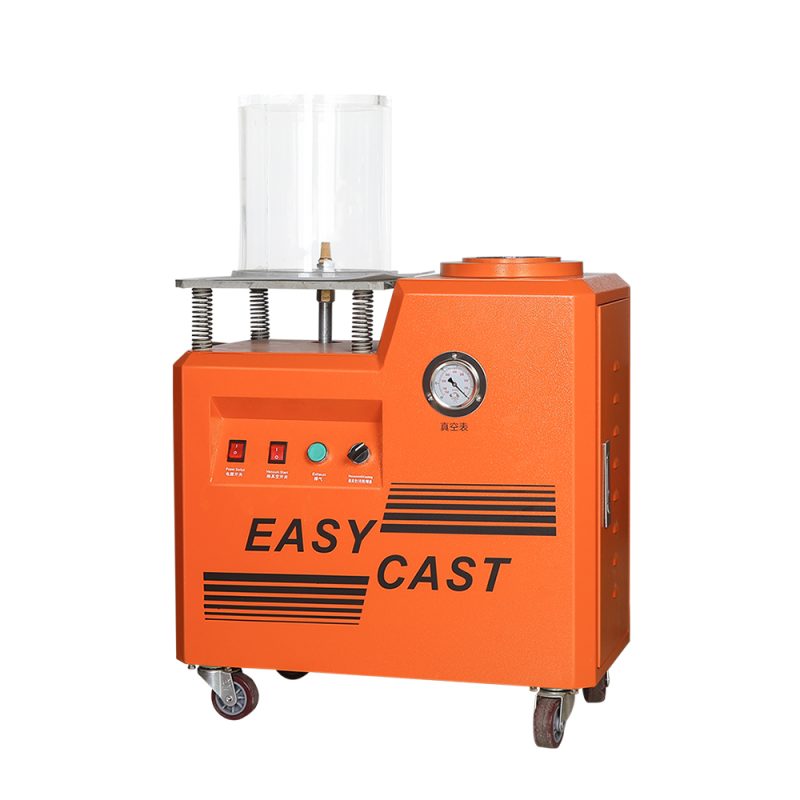
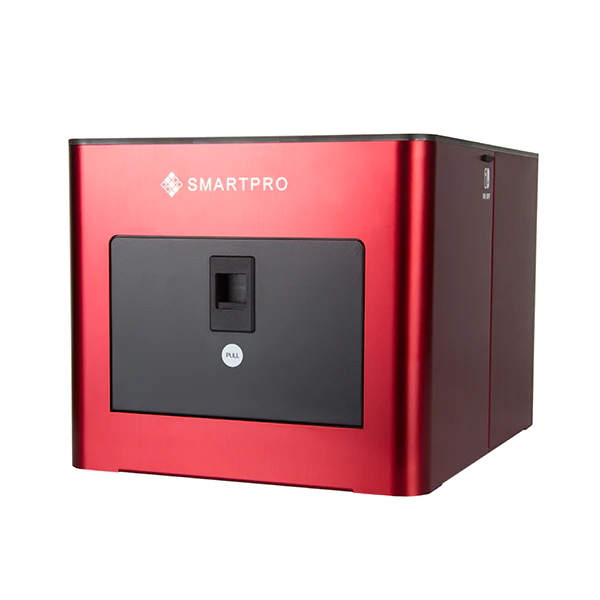
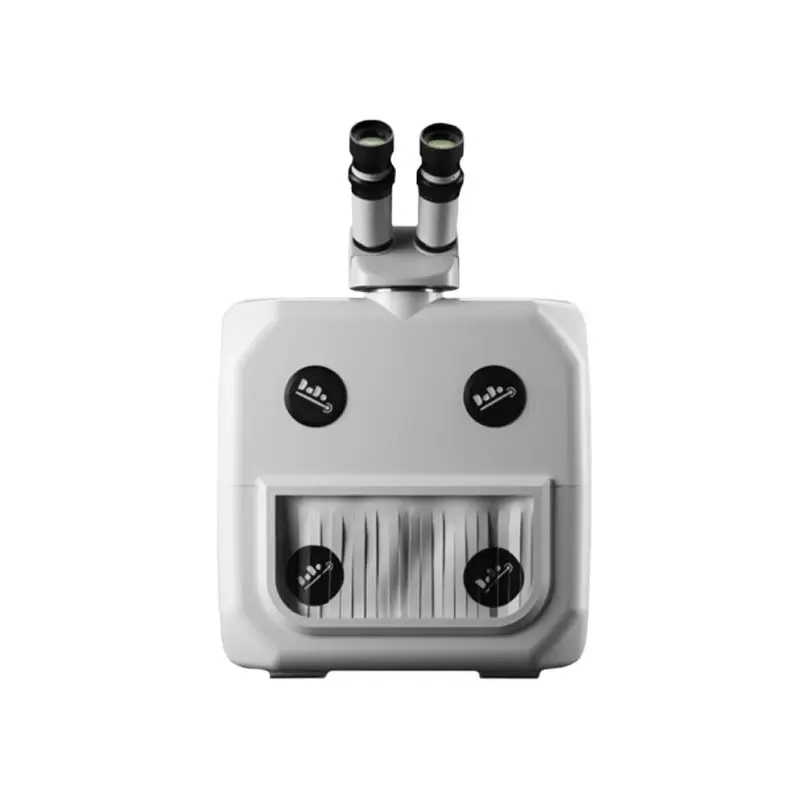
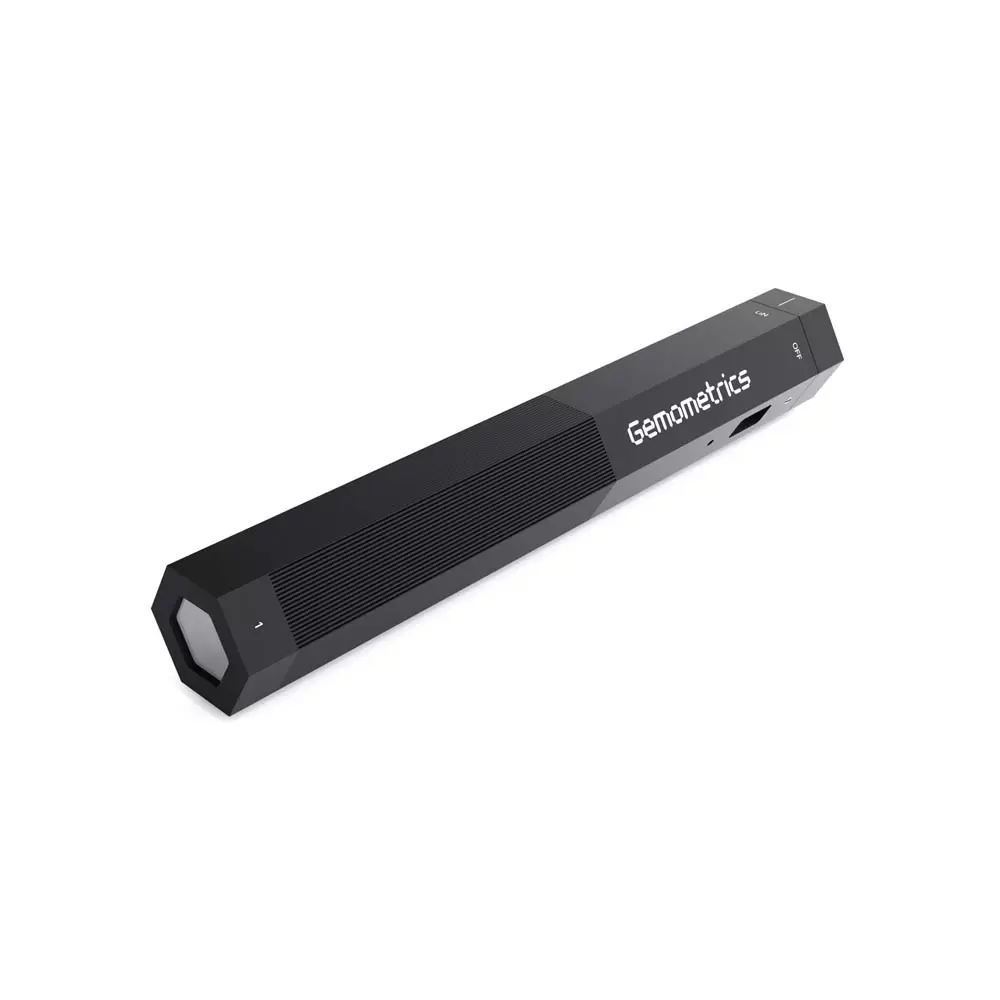
के तपाईलाई सुन परीक्षण मेसिन चाहिन्छ

Shipping Perks
Free standard shipping
within Kathmandu.
Return Policy
Full Return with reason if not satisfied.

Customer Service
Call us if you have any questions.

Genuine Product
We provide genuine products.
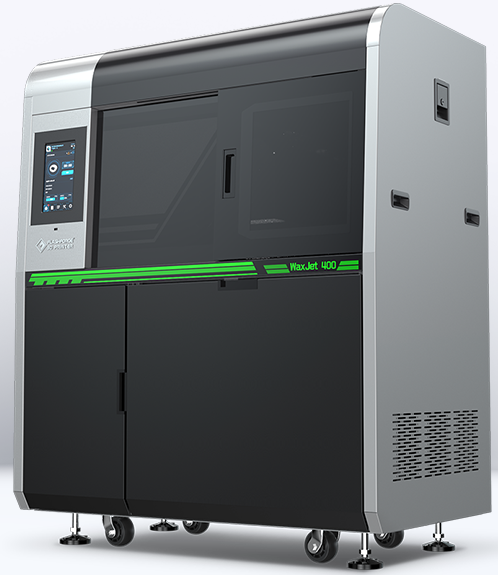
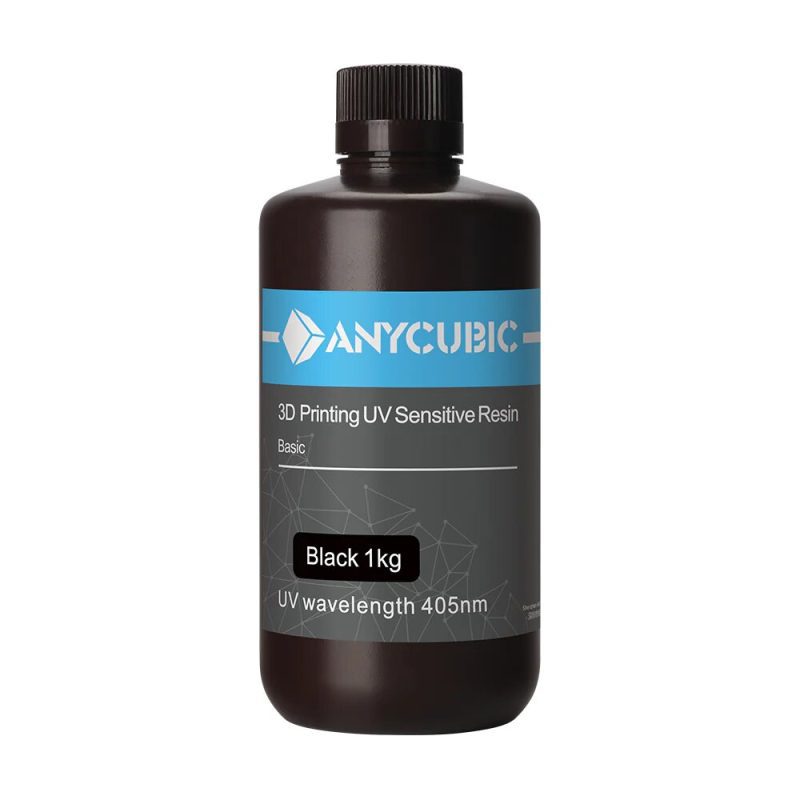
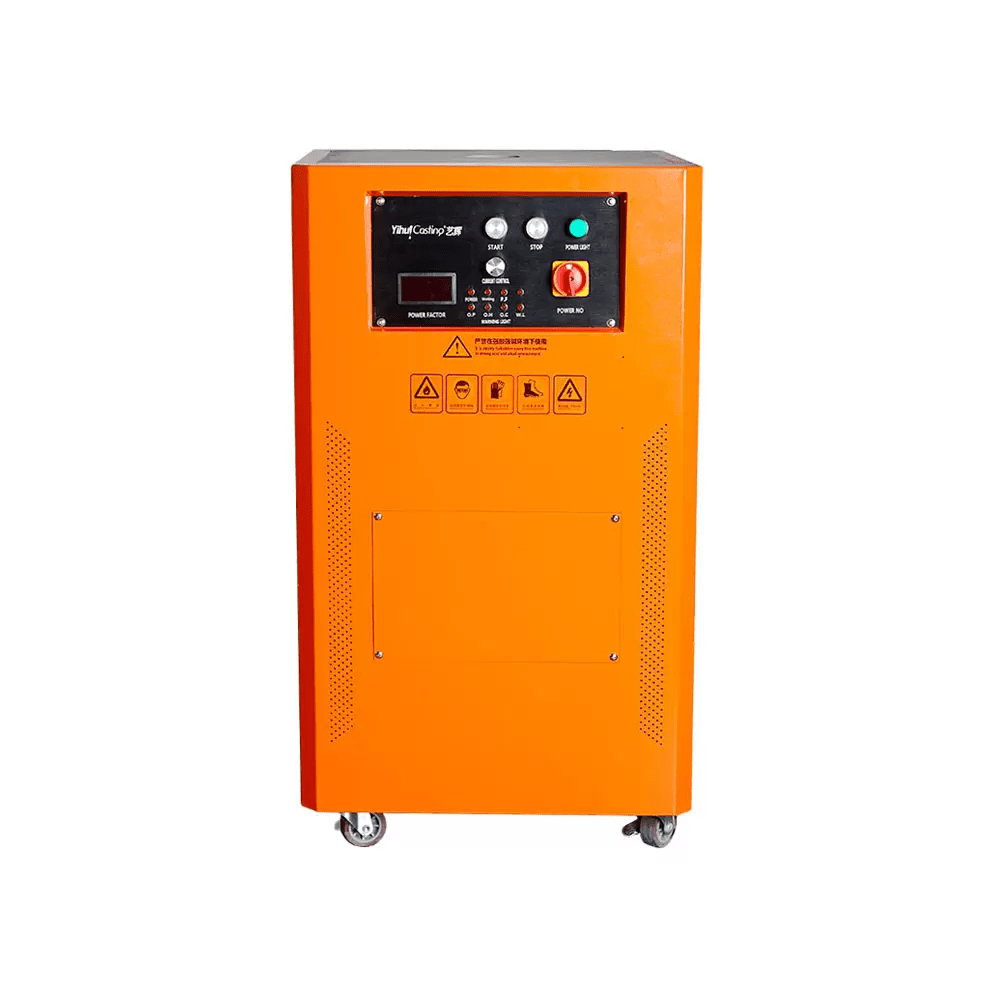
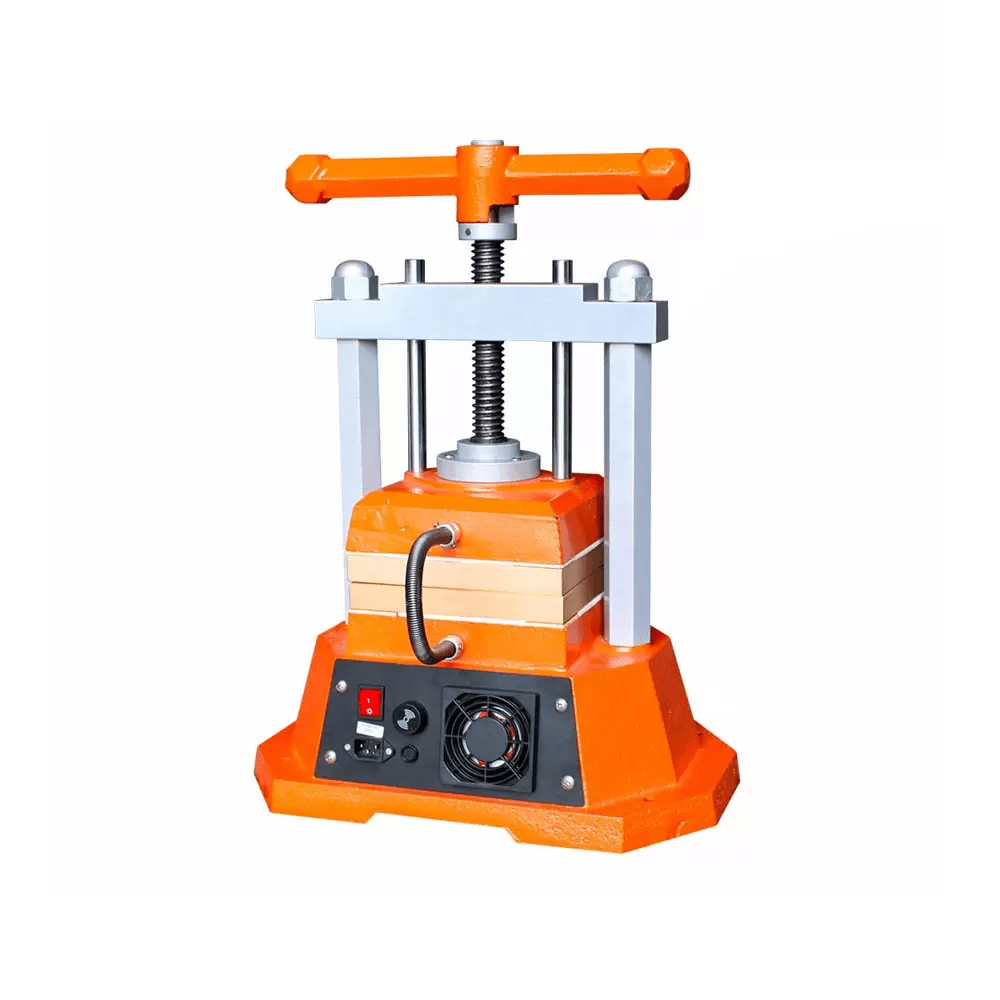
TOP SALE ON THIS WEEK
Features Products
OUR BUSINESS PARTNERS